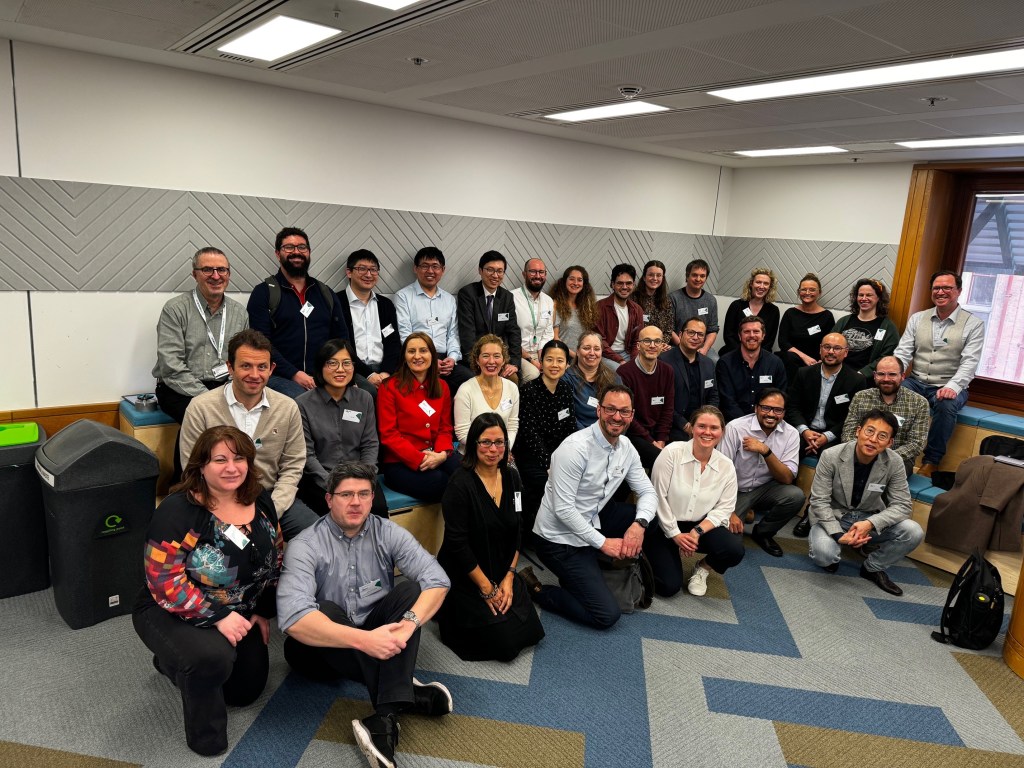
In this blog entry we discuss the outcomes of the recently held JavaScript for Visualisation Workshop, The Genome Analysis Centre (TGAC), Norwich, UK, April 21-22. This workshop brought 26 attendees from around the UK and the continent. The event was open to both core technical members and developers not currently involved in the BioJavaScript (BioJS) project. The workshop was facilitated by Software Sustainability Institute (SSI) 2016 fellow Manuel Corpas to promote software best practices and the latest standards in JavaScript, inspiring the future technical roadmap for the BioJS community.

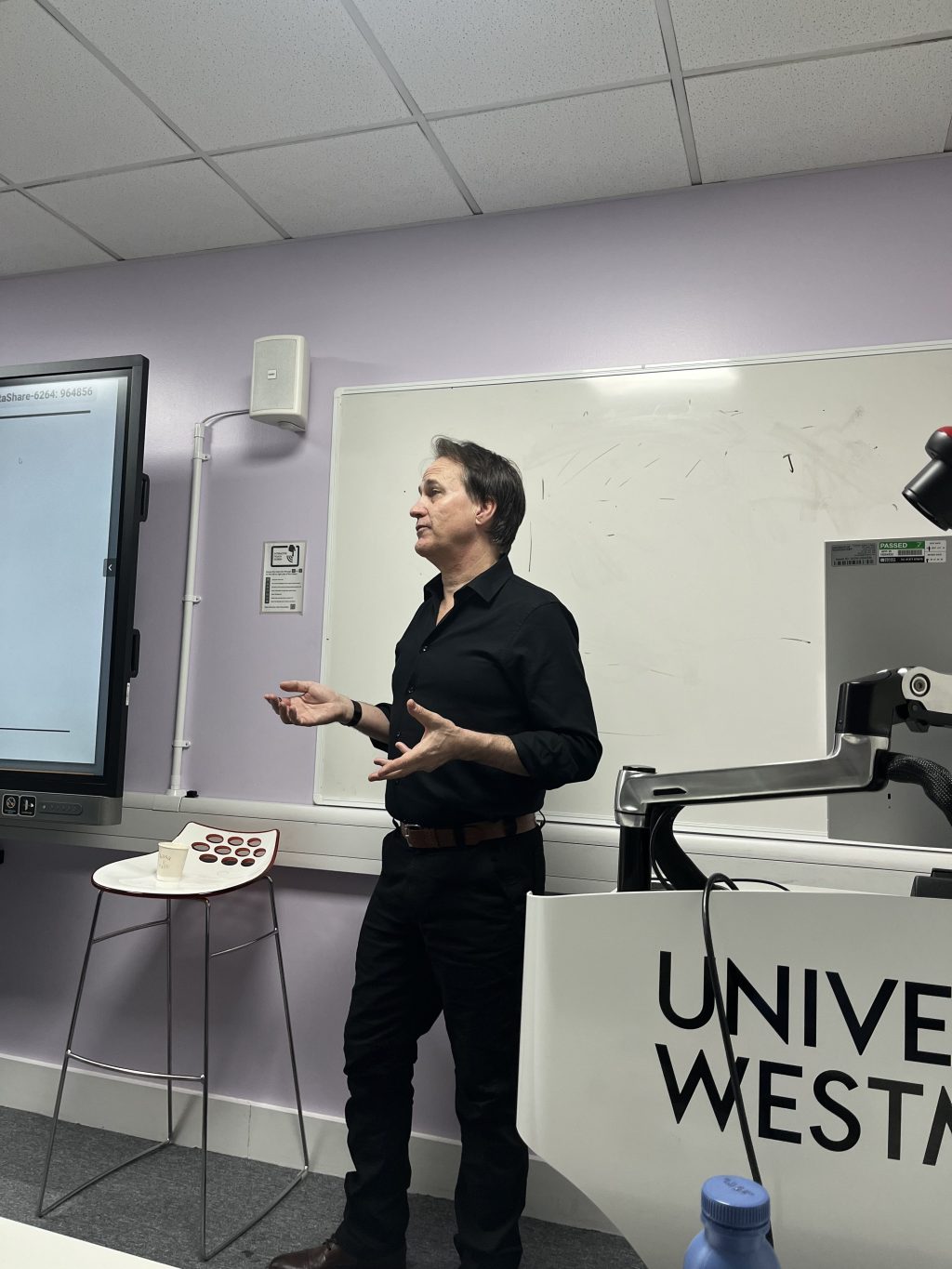
Tim Ruffles teaching at TGAC’s training suite during the workshop
Keywords: JavaScript, BioJS, Software Best Practices, Web Components, D3, Bioinformatics
-
The Need for this Workshop
JavaScript has become the de facto language of the web. Console-like interactivity is now common place in most web applications and sites. Online collaboration plays an important role in research, enabling scientists to share and disseminate results regardless of geographical location. The data-intensive field of bioinformatics greatly benefits from JavaScript technologies applied to visualisation of biological datasets. The JavaScript Visualisation Workshop held at TGAC on April 21-22 brought together scientific programmers and developers from around the UK and Europe to Norwich to learn about the latest cutting-edge methodologies and concepts applicable to biological web visualisation. Hosted under as an outreach event for the project, this workshop provided compelling guidelines for good practice development in BioJS and the wider bioinformatics community as a whole.
2. Contributions from SSI 2016 Fellow Manuel Corpas
Corpas was the leading facilitator of the course syllabus, promoter of the event via social media and recruiter of senior JavaScript developer and consultant Tim Ruffles. Corpas was supported by TGAC’s Marketing and Communication and TGAC’s Scientific Training & Education Programme to target the wider bioinformatics community via social media and the organisation of the logistics on-site. Corpas also engaged both the BioJS community and Ruffles through a number of online discussions when designing the syllabus for the workshop.
3. Why We Chose an Outside Expert
The latest technological developments for any programming language applied to biology are not originated by bioinformaticians but software engineers. Industrial standards for production-ready software are thus set by experts who work full time on the language development. It is thus imperative for bioinformaticians, the target audience of this workshop, to access first hand the latest technological trends for JavaScript from mainstream contributors. Senior JavaScript developer and mentoring coach, Tim Ruffles, led the workshop, providing both technical introductions and hands-on exercises on modern ES6 syntax, web components and NPM development. Tim has a strong reputation in the UK as a JavaScript contractor, trainer and Google Developer Expert for AngularJS, a development framework. The photograph below shows Tim explaining ES6 to a full-house Darwin training suite at TGAC.
4. How this Workshop Will Shape the BioJS Community
During the second day of the workshop, emphasis was placed on the basics of BioJS component development. This meant that attendees who were not familiar with BioJS had a chance to experiment with it. The philosophy of BioJS implies development of self-contained components that can be reused, extended and combined via an open source license like Apache 2 or MIT. In the words of Antoniya Stavreva, who travelled from Bulgaria to attend the workshop, “[she] learnt how to develop and extend BioJS components more efficiently, making it possible for other people to reuse them.” Overall, this workshop enabled the advancement of best practices in software engineering for 26 attendees, many of them not familiar with the BioJS project, yet exposing them to best practices on how to develop reusable components. This training also had a tremendous impact on core BioJS developers, who enhanced their expertise on crucial technologies that will affect the future of the project.
Acknowledgements
I would like to acknowledge the contribution of material for this post from Jessica Jordan, Carlos Horro, Haydee Artaza and Antoniya Strateva. I am particularly grateful to TGAC’s, Scientific Training & Education Programme who supported and managed all the logistics for this workshop: Emily Angiolini, Matt Drew, Helen Tunney and Paul Yorke. I am also grateful to Chris Bennett and Sasha Stanbridge from the TGAC Marketing and Communications team for their management of the social media campaign, the design of the workshop banner and for taking pictures.













































Leave a comment